What if pretrained weights are a genome, not a brain?
Experiments testing whether pretrained weights encode structural blueprints rather than knowledge, and what that means for how we build AI.
I've been running experiments for the past few weeks that started from a simple question and led somewhere I didn't expect. I want to share what I found, what it might mean, and where the gaps are. None of this is a claim. It's an observation with data behind it. I could be completely wrong.
How this started
I was thinking about evolution and backpropagation. Not the mechanics of either, but what they have in common at a higher level. Both are search algorithms. Both find structure in the data they operate on. And when you press different search algorithms against the same data, they converge on the same structure. CNNs and Vision Transformers are architecturally alien to each other, but their learned representations are remarkably similar. The searcher is interchangeable. The data is the message.
Then I ran a thought experiment. Imagine evolution, but instead of running over billions of years against a constantly changing reality, it ran at a single instant. One frozen snapshot of reality. No variation over time. No predator-prey arms races. No changing seasons. What kind of intelligent solution would it produce?
The answer: a crystal. A perfect, dense, hierarchical map of that frozen instant. No flexibility needed because nothing changes. No memory because there's nothing to track over time. No prediction because nothing will be different tomorrow. Just structure mirroring structure, at every scale, perfectly fitted and completely rigid.
And then it hit me. That's what LLMs are. Trained on a frozen corpus. Frozen weights at deployment. Every regularity mapped at high resolution. The crystal.
That's what LLMs are. Trained on a frozen corpus. Frozen weights at deployment. The crystal.Three stages, not two
This realization opens a bigger question. Evolution didn't just produce a crystal. It produced something far more interesting. And the difference isn't about optimization or scale. It's about stages.
Stage 1: Evolution produces the genome. A search over deep time that finds compressed structural biases. The output is small (3 billion base pairs in humans) relative to the search space it explored. This is pretraining. Backprop finds biases in the training data and compresses them into weights.
Stage 2: The genome produces the brain. This is development. The genome doesn't specify every synapse. It specifies wiring rules, density patterns, connectivity principles. Over five years, these rules interact with sensory experience to build a cognitive architecture. Infant amnesia. And here's the remarkable part: 99% of five-year-olds have the same cognitive machinery regardless of whether they were born in a Pacific island jungle in 3000 BC or San Francisco in 2025. Same object permanence. Same causal reasoning. Same theory of mind. Different knowledge. Same architecture. Universal output from any sufficiently rich input.
Stage 3: The brain learns. The architecture that development built now acquires knowledge and skill. This runs on 20 watts. It's sample efficient. It resists catastrophic forgetting. It can do novel reasoning that no training example demonstrated.
Current AI does Stage 1 and jumps straight to Stage 3. We pretrain (evolution), then fine-tune and deploy (try to make it learn and reason). We completely skip Stage 2. And maybe that's why we need 100 megawatts instead of 20.
I don't know if this framing is correct. But I wanted to test whether Stage 1 actually produces something genome-like. Not the brain. Not the final intelligence. The blueprint.
Current AI does Stage 1 and jumps straight to Stage 3. We completely skip Stage 2. And maybe that's why we need 100 megawatts instead of 20.The skeleton experiment
If the weights are a genome, then the structural information should be separable from the specific weight values. Think of it this way: the genome doesn't specify the exact strength of every synapse. It specifies which connections should exist. The wiring diagram.
So I took a pretrained language model (SmolLM2-135M), looked at which weights were large by magnitude, and created a binary mask: 1 where the weight was in the top 10% by magnitude, 0 everywhere else. That mask is the skeleton. The connectivity pattern.
Then I created a brand new randomly initialized model with the same architecture. Applied the pretrained skeleton to it. Zeroed out weights where the mask said 0. Enforced the mask after every training step so the dead connections stay dead. The model can only learn in the positions where the pretrained model had large weights.
For comparison: another fresh random model, same architecture, same sparsity (90% of weights zeroed), but the mask is completely random. Same number of active connections, but the pattern is arbitrary.
Both models were trained on FineWeb-Edu for 500 steps. Same data, same optimizer, same everything. The only difference is the pattern of which connections are allowed to learn.
The sparsity sweep
I ran this at every sparsity level from 10% to 90%.
Skeleton value scales exponentially with sparsity
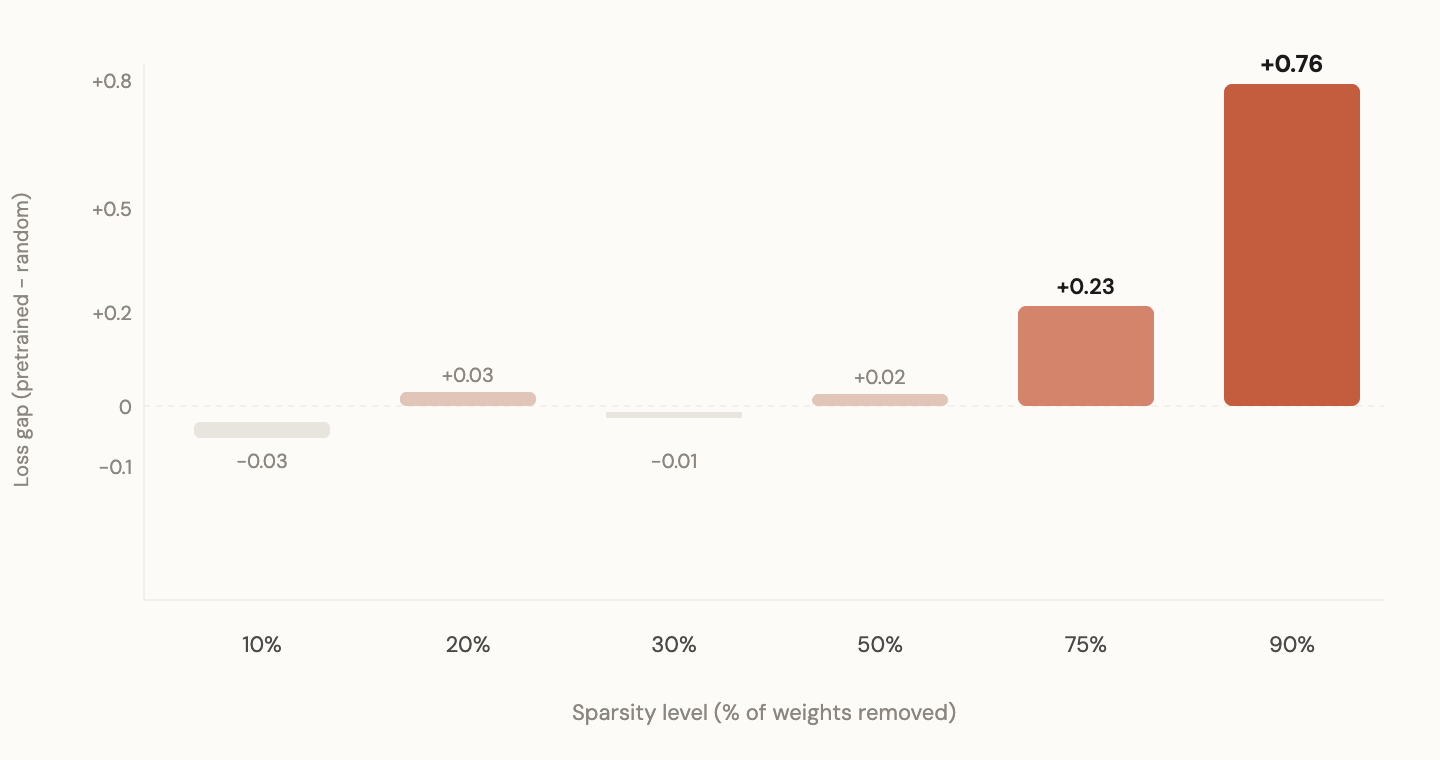
At 10% sparsity (barely any weights removed), both patterns perform identically. At 30%, still no difference. At 50%, a faint hint. At 75%, a clear gap. At 90%, an enormous gap.
The pattern's value scales exponentially with sparsity. When you have plenty of connections, it doesn't matter where you put them. When resources are scarce, knowing where to allocate them becomes everything.
This makes sense through the genome lens. The human genome's value is also exponential under resource constraints. Brains can't be fully connected. Bodies can't maintain infinite synapses. The genome is the answer to the question: when you can't afford everything, what do you build?
When resources are scarce, knowing where to allocate them becomes everything.Cross-modal: language skeleton on vision
If the skeleton encodes language-specific structure, it should only help on language. If it encodes something more universal about how transformers organize computation, it might help on other tasks too.
Cross-domain: language skeleton helps vision at every scale
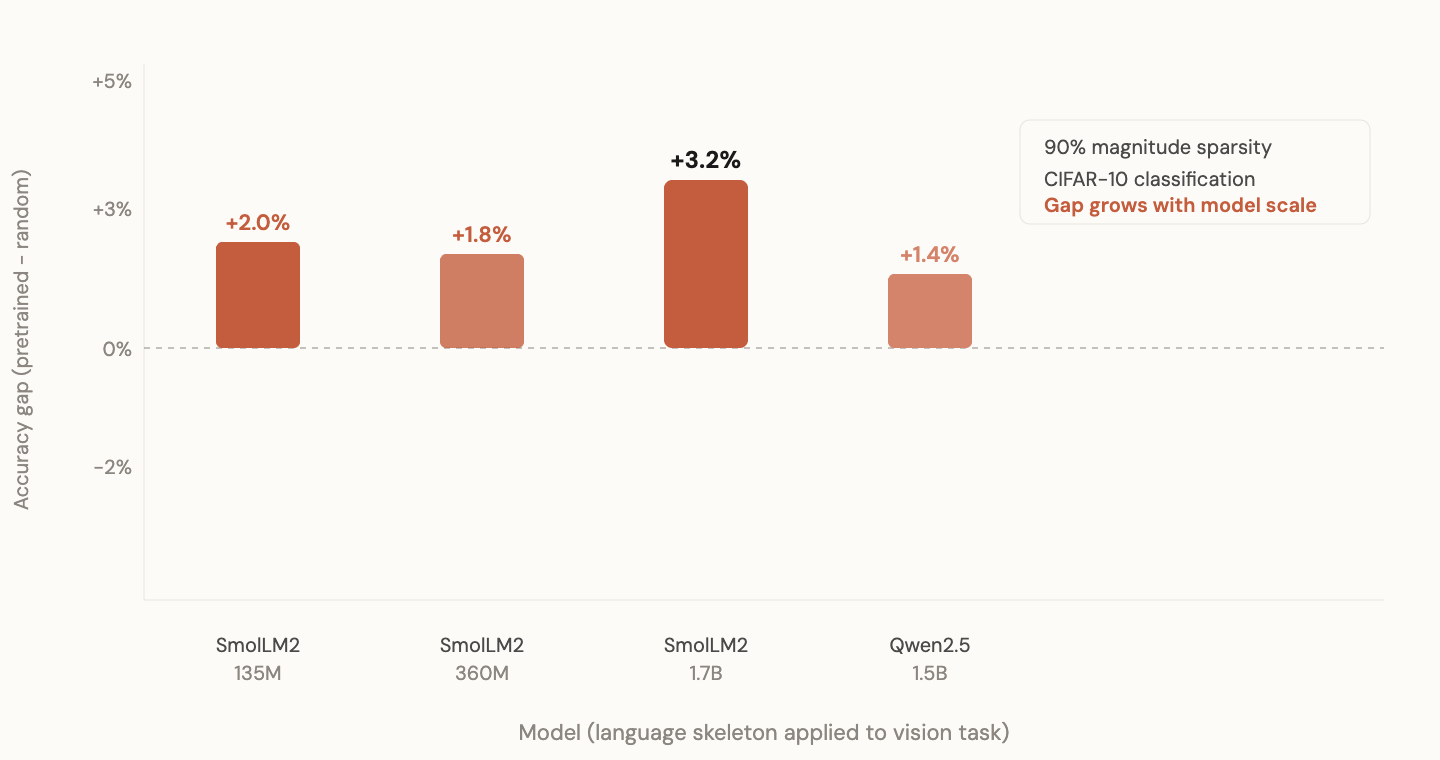
I took the language skeleton from SmolLM2-135M and tested it on CIFAR-10 image classification. Same architecture, but the input embedding and output head are swapped for vision (pixel patches in, class labels out). The transformer body and its skeleton are unchanged.
Three seeds: 42, 123, 7. The language skeleton won on all three. Mean accuracy gap: +2.5%. Small but consistent. Never flipped sign across seeds.
At 1.7B scale (SmolLM2-1.7B), the vision gap actually grew to +3.2%. Larger models encode more universal structural information.
| Seed | Lang Skeleton | Rand Skeleton | Gap |
|---|---|---|---|
| 42 | 53.1% | 50.8% | +2.4% |
| 123 | 54.9% | 53.0% | +1.9% |
| 7 | 55.5% | 52.3% | +3.1% |
| Mean | 54.5% | 52.0% | +2.5% |
Magnitude breaks. SVD doesn't.
Here's where it got interesting. I tried the same magnitude-based skeleton experiment on Qwen 2.5-1.5B. It failed. The random skeleton actually won on language by 0.53 nats. Complete reversal.
For a while I thought the whole thing was architecture-specific or coincidental. But then I thought about what magnitude actually measures. When you pick the largest weights, you're looking at individual entries. One number in a matrix, in isolation. But a weight matrix is a transformation. It has structure that lives in the relationships between weights, not in any single weight.
SVD recovers structural signal where magnitude fails
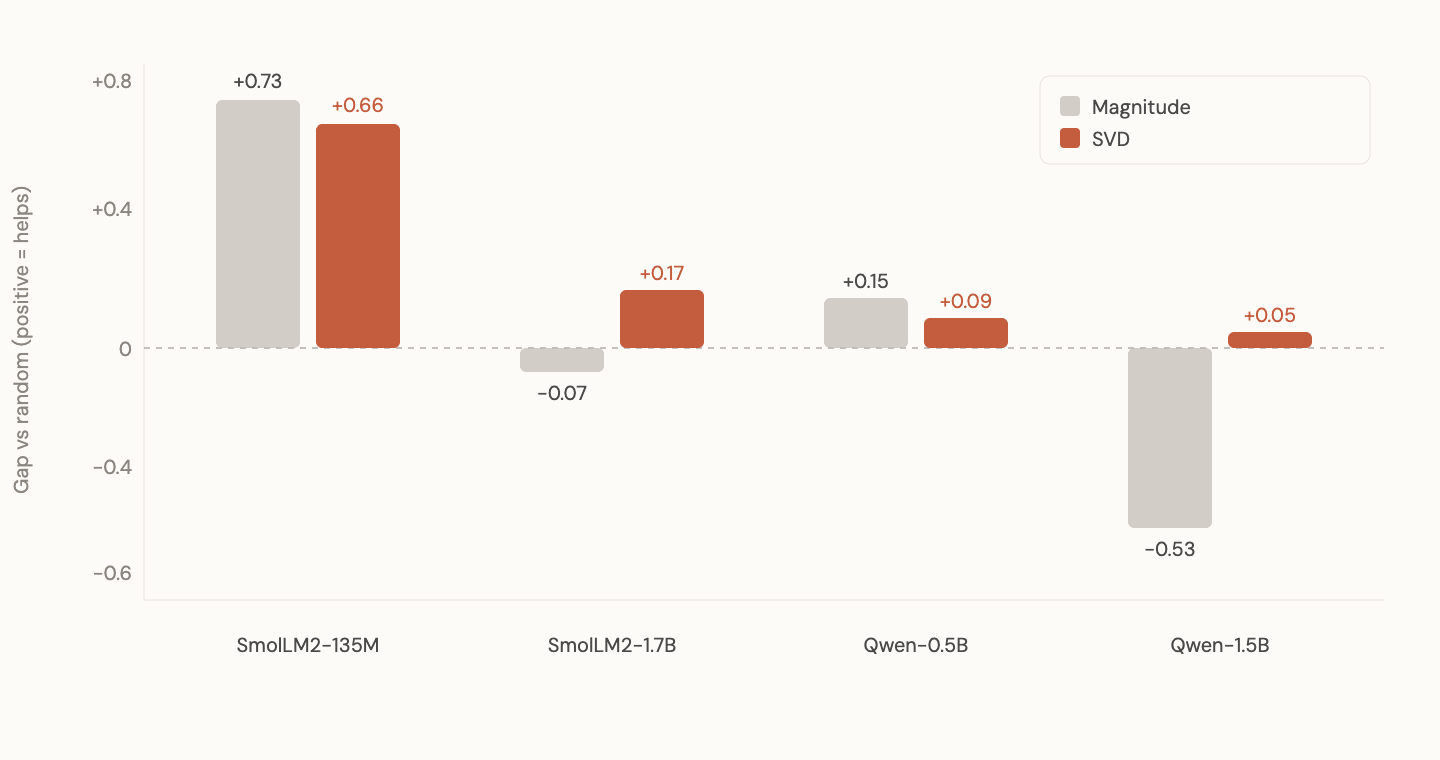
SVD (Singular Value Decomposition) decomposes a matrix into its principal directions. The directions along which the transformation does real work. Instead of asking "which individual weights are large," SVD asks "which directions does this matrix use?"
I extracted the top 10% of singular directions from each model and used those to initialize fresh random models (random magnitudes, but constrained to the pretrained subspace).
The results across all four model families (SmolLM2-135M, SmolLM2-1.7B, Qwen 2.5-0.5B, Qwen 2.5-1.5B): every single one positive. SVD recovered signal where magnitude failed. On Qwen 1.5B specifically, the result flipped from -0.53 (magnitude) to +0.05 (SVD).
The structural information is in the subspace geometry, not in individual weight magnitudes. Magnitude is a crude proxy that works on small models and breaks at scale. SVD reads the actual structure.
The structural information is in the subspace geometry, not in individual weight magnitudes.The convergence result
This is the experiment I'm most confident about.
I took four Pythia-160M models trained by EleutherAI. Same architecture. Same training data (the Pile). Same training order. Different random seeds. Four completely independent training runs that started from different random points and ended at different weight matrices.
Four seeds, one geometry (functional convergence)
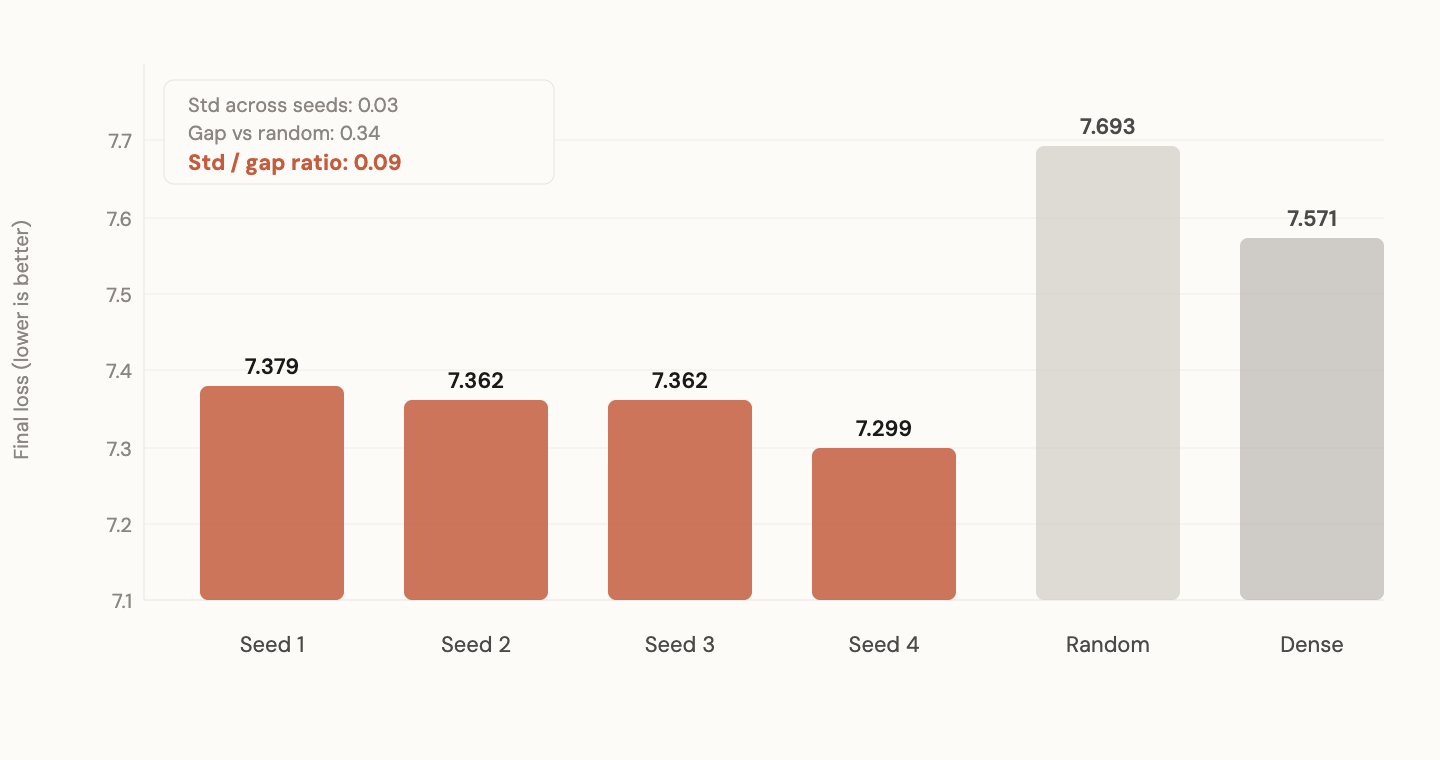
I extracted SVD axes (top 10% singular directions) from each. Gave each set of axes to a fresh model with random magnitudes. Trained all four on FineWeb-Edu for 200 steps. Also trained a random-axes baseline and a dense (unconstrained) baseline.
The results:
| Condition | Final loss |
|---|---|
| Seed 1 axes | 7.378 |
| Seed 2 axes | 7.362 |
| Seed 3 axes | 7.362 |
| Seed 4 axes | 7.299 |
| Random axes | 7.693 |
| Dense (all directions) | 7.571 |
Standard deviation across seeds: 0.03. Mean gap over random: 0.34. The ratio (std/gap) is 0.09.
Four independent training runs found the same computational subspaces. The initialization doesn't matter. The architecture plus data forces the solution geometry. The specific weights differ. The directions don't.
And notice: 10% subspace beats 100% dense in early training. A model that can only move in the directions that training always finds outperforms a fully unconstrained model. Less is more, if you know where to look. And all four seeds know the same "where."
10% subspace beats 100% dense in early training. Less is more, at least at the beggining if you know where to look. And all four seeds know the same "where."The permanent gap
I trained the Pythia models to convergence (~40K steps). Dense eventually overtakes the 10% pretrained subspace, which is expected since it has 10x the capacity. But random 10% never catches pretrained 10%. Even with unlimited training, the gap between pretrained and random subspace does not close.
The permanent gap: random subspace never catches up
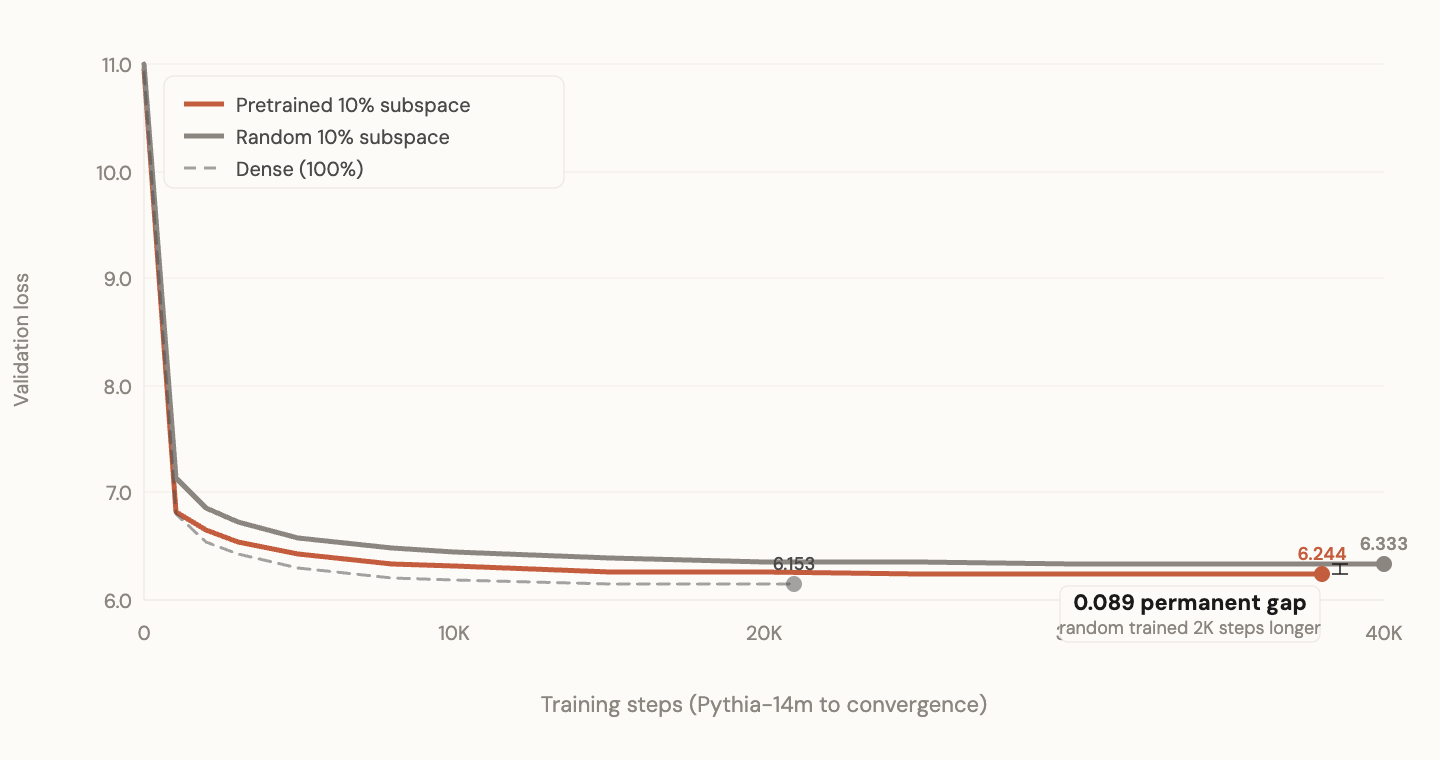
Zoomed: the gap at convergence never closes
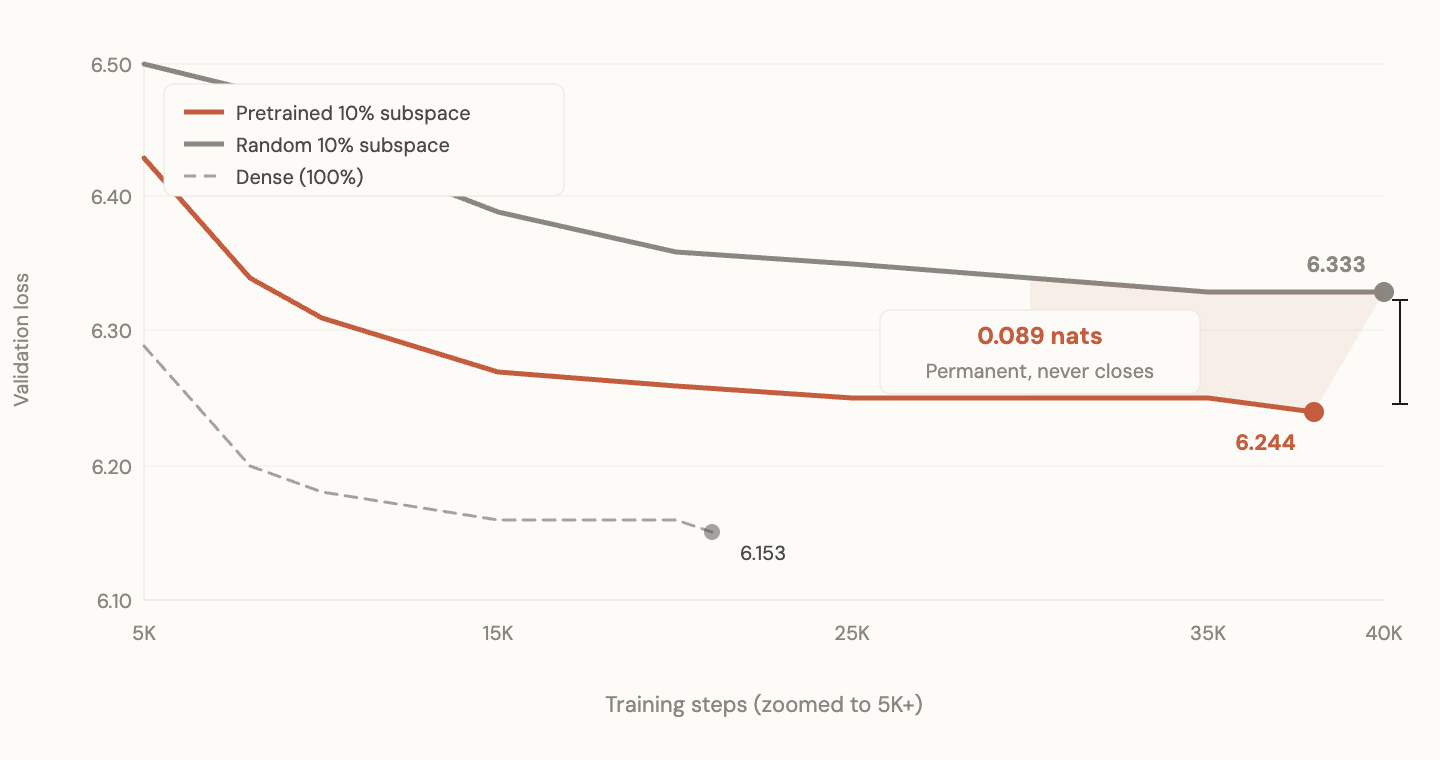
This is the result that matters most.
If pretrained subspace just helped learn faster, random would eventually catch up. It doesn't. The pretrained directions give access to a region of loss landscape that random directions cannot reach, regardless of how long you train. The subspace defines what solutions are reachable. Wrong directions mean some solutions are permanently inaccessible under resource constraints.
The genome doesn't make brains learn faster. It makes certain kinds of learning possible that would otherwise be impossible in a resource-constrained system.
The genome doesn't make brains learn faster. It makes certain kinds of learning possible that would otherwise be impossible in a resource-constrained system.What does this mean
I'm going to be careful here because I've been wrong before and I have a strong prior toward this framing that I can't fully separate from confirmation bias.
What I think the data shows:
-
Pretrained models contain structural information (which subspaces to use) that is separable from the specific weight values.
-
This structural information is deterministic. Given an architecture and sufficiently rich data, training always converges to the same computational directions, regardless of initialization.
-
The information partially transfers across modalities. Language-derived structure helps vision. The signal is small but consistent.
-
Magnitude-based extraction works for small models but breaks at scale. SVD-based extraction works universally. The information lives in subspace geometry, not individual weight magnitudes.
-
The structural information defines what is learnable under resource constraints. It doesn't just speed up learning. It makes some learning permanently possible that would otherwise be permanently impossible.
What I think this suggests but cannot yet prove:
Current pretraining is doing something analogous to evolution's outer loop. It's finding structural biases. Not knowledge, not skills, not intelligence. The blueprint for a computational architecture. And we're currently deploying the blueprint as if it were the final product. We're trying to make the genome reason, instead of using the genome to build something that reasons.
The developmental stage (Stage 2) is missing from current AI. Building it would mean taking the structural information from pretraining and using it to construct a separate, small, efficient learner that operates in its own parameters. Not fine-tuning. Not distillation in the current sense. Literally constructing a new network whose architecture is determined by the pretrained model's structural fingerprint.
I don't know how to do that yet. But I think the experiments show that the structural fingerprint exists, that it's universal, and that it matters.
What I got wrong along the way
The Qwen magnitude failure was important. I almost concluded the whole thing was architecture-specific before realizing magnitude was the wrong extraction method. If I'd stopped there, I would have missed the SVD insight.
The vision experiments were messy. Causal attention on image patches is a terrible way to do vision. The dense baseline on CIFAR-10 was confounded by regularization effects of sparsity on a tiny dataset. I ran too many variations before controlling properly.
I spent too long on cross-modal transfer before doing the convergence experiment, which turned out to be the cleanest and most important result.
What's next
I don't know. The boundary conditions I haven't tested include: whether the convergence holds across different training data (not just different seeds), whether there's a clean way to test cross-modal SVD transfer, whether the structural information is in the per-layer rank profile or in the specific singular vectors, and whether any of this can be turned into an actual Stage 2 system.
If you see a flaw in the experiments or the reasoning, I genuinely want to hear it. I'm one person running these on a single GPU and I know my bias toward this framing is strong. The experiments are designed to test the framing, not confirm it. Several of them returned null or negative results and I've reported all of them.
If you see specific flaws, reach out. All raw results including the failures are in the results log.
Keep reading
April 1, 2026·10 min read
One neuron changes everything: the 63 vs 64 puzzle
A 3-layer MLP learning F = m*a behaves completely differently with 63 neurons vs 64. The investigation reveals three training regimes, winner-take-all gradient dynamics, and a lesson about what you're actually ablating when you change architecture.
March 17, 2026·16 min read
The genome hypothesis: a thought experiment about what pretraining actually finds
A constraint-based argument for why pretraining might be evolution's outer loop, what the missing developmental stage looks like, and what that means for how we build AI.
April 1, 2026·9 min read
Grokking has two phases, and you can see the boundary
A component-freezing ablation study on modular division reveals that grokking isn't one process. It's two: infrastructure setup, then computational reorganization. The boundary between them is sharp and measurable.